
This web site, https://www.vcru.wisc.edu/sdata, contains data from the laboratory of Philipp W. Simon, USDA-ARS Vegetable Crops Research Unit [Click here for our web page]
 |
USDA ARS VCRU Data Server This web site, https://www.vcru.wisc.edu/sdata, contains data from the laboratory of Philipp W. Simon, USDA-ARS Vegetable Crops Research Unit [Click here for our web page] |
This carrot linkage map was published in:
Microsatellite isolation and marker development in carrot -
genomic distribution, linkage mapping, genetic diversity analysis and marker transferability
across Apiaceae
Pablo F Cavagnaro, Sang-Min Chung, Sylvie Manin, Mehtap Yildiz, Aamir Ali, Maria S Alessandro,
Massimo Iorizzo, Douglas A Senalik and Philipp W Simon
BMC Genomics 2011, 12:386 doi:10.1186/1471-2164-12-386
Background
The Apiaceae family includes several vegetable and spice crop species among which carrot is the most
economically important member, with ~21 million tons produced yearly worldwide. Despite its importance,
molecular resources in this species are relatively underdeveloped. The availability of informative, polymorphic,
and robust PCR-based markers, such as microsatellites (or SSRs), will facilitate genetics and breeding of carrot
and other Apiaceae, including integration of linkage maps, tagging of phenotypic traits and assisting positional
gene cloning. Thus, with the purpose of isolating carrot microsatellites, two different strategies were used; a
hybridization-based library enrichment for SSRs, and bioinformatic mining of SSRs in BAC-end sequence and EST
sequence databases. This work reports on the development of 300 carrot SSR markers and their characterization at
various levels.
Results
Evaluation of microsatellites isolated from both DNA sources in subsets of 7 carrot F2 mapping populations
revealed that SSRs from the hybridization-based method were longer, had more repeat units and were more
polymorphic than SSRs isolated by sequence search. Overall, 196 SSRs (65.1%) were polymorphic in at least one
mapping population, and the percentage of polymophic SSRs across F2 populations ranged from 17.8 to 24.7.
Polymorphic markers in one family were evaluated in the entire F2, allowing the genetic mapping of 55 SSRs (38
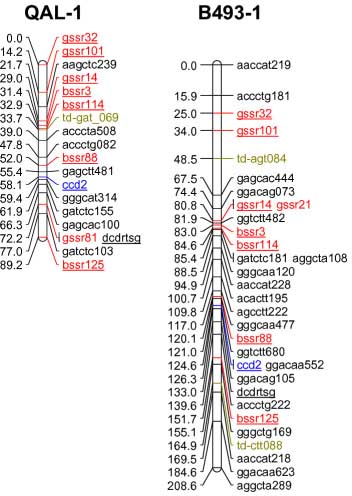
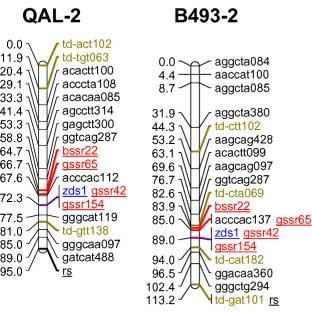
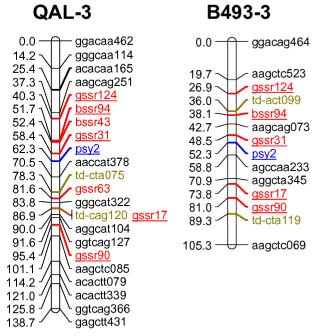
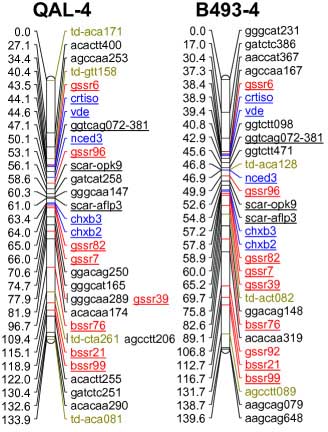
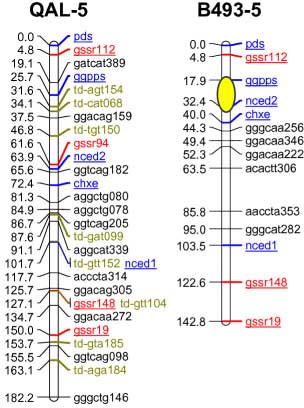
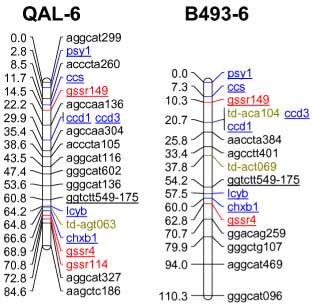
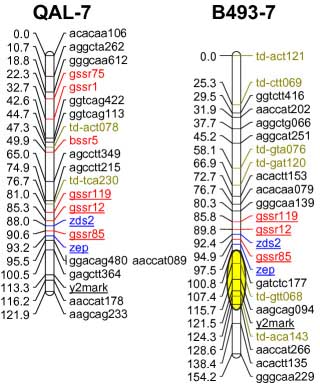
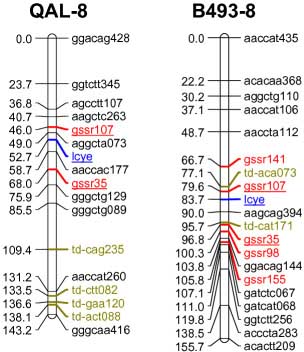
codominant) onto the carrot reference map. The SSR loci were distributed throughout all 9 carrot linkage groups
(LGs), with 2 to 9 SSRs/LG. In addition, SSR evaluations in carrot-related taxa indicated that a significant
fraction of the carrot SSRs transfer successfully across Apiaceae, with heterologous amplification success rate
decreasing with the target-species evolutionary distance from carrot. SSR diversity evaluated in a collection of
65 D. carota accessions revealed a high level of polymorphism for these selected loci, with an average of 19
alleles/locus and 0.84 expected heterozygosity.
Conclusions
The addition of 55 SSRs to the carrot map, together with marker characterizations in six other mapping
populations, will facilitate future comparative mapping studies and integration of carrot maps. The markers
developed herein will be a valuable resource for assisting breeding, genetic, diversity, and genomic studies of
carrot and other Apiaceae.
This population was earlier referenced by Carlos Santos (v.1), Brian Just (v.2), and Dariusz Grzebelus (v.3)
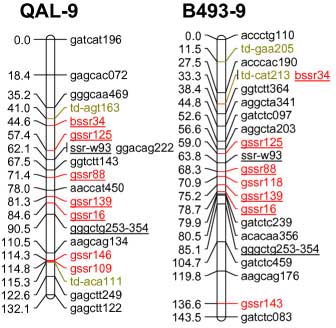
| Marker info | Left=QAL Right=B493 | Marker info |
|---|---|---|
 |
||
 |
||
 |
||
 |
||
 |
||
 |
||
 |
||
 |
||
 |