================================
Parallel computing with BAPS 5
================================
- Jing Tang 22/11/2007
===============================
Q1: What does parallel mean here?
===============================
A1: Parallel computing aims to
distribute the tasks of BAPS analysis into multiple computers such that the
results will be produced faster. The idea is based on the fact that the BAPS
analysis usually can be divided into smaller parallel tasks, which may be
carried out simultaneously.
===============================================================
Q2: In what kind of a situation can
I run parallel computing?
===============================================================
A2: Parallel computing is enabled
for both the mixture and admixture analysis, but with different purposes.
A2.1 Mixture analysis
For mixture
analysis, parallel computing can make significant improvement on decreasing the
computation time if the data is large and fuzzy. The fuzziness of a dataset is
evaluated by the uncertainty of finding the optimal partition in limited runs.
Of course one cannot know the fuzziness of the data prior to the analysis, but
for large datasets it is always recommended to use multiple estimation runs in
BAPS analysis by choosing different initial number of clusters, and to check
whether the partition result from each run is stable or not.
The idea of
parallel computing is to distribute the task of multiple runs into multiple
computers, such that on each computer only a subset of the possible initial
cluster numbers are tested. The partition result identified by each 'local'
computer will be compared with each other such that the 'global' optimal
partition can be determined.
A2.2 Admixture analysis
The parallel
computing for admixture analysis is based on the algorithm in which the
individuals of one cluster in a partition can be computed independently from
other clusters. Thus each cluster can be made to run on a separate computer to
achieve the results faster.
Therefore,
parallel computing for admixture is always recommended for large and complex
data sets as long as there are multiple computers available.
A2.3 The case where parallel
computing should NOT be used
IF LINKAGE
CLUSTERING model is used WHEN the data contains missing values, and the data
HAS NOT been pre-processed into a .mat format, then parallel computing is not
allowed. For such a case, missing values are estimated stochastically during
the pre-processing step. The complete data used afterwards in the mixture analysis
might not be identical for different runs. This data inconsistency will
invalidate the ground for parallel computing since the partition results are no
longer comparable. However, parallel computing can be used if the data with
missing values has already been pre-processed. For the independent loci model
the parallel computing is ALWAYS allowed.
============================================
Q3: How to carry out the parallel
computing?
============================================
A3: Suppose we want to compare the
partition results given by different initial number K of clusters. Such a task
can be distributed by parallel computation into multiple computers, each of
which handles only a subset of Ks. For instance we have a dataset called
'simple_data.txt' and we need to compare the partition results based on initial
K = 2, 3, 4, 5. Given that two computers are available, the whole process will
be implemented in the following:
1) Write a script file that contains
the parameters to run a subtask in the first computer. Here we give a script
example indicating that mixture analysis should start with K=2 and 3 (Figure
1). The details of writing a script are described in section Q4.
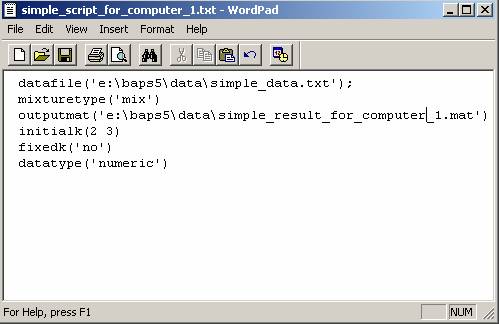
Figure 1: A script prepared for a mixture analysis on
computer No. 1. It starts a mixture analysis with initial K=2 and 3.
2) Open a command prompt window if
using Windows, or open a shell window if using Linux/Unix. Go to the directory
where the BAPS package is installed. Type 'baps5' followed by the COMPLETE name
of the script file and press ENTER. NB! Change the string ‘baps5’ in the
command according to the program version used. (Figure 2)
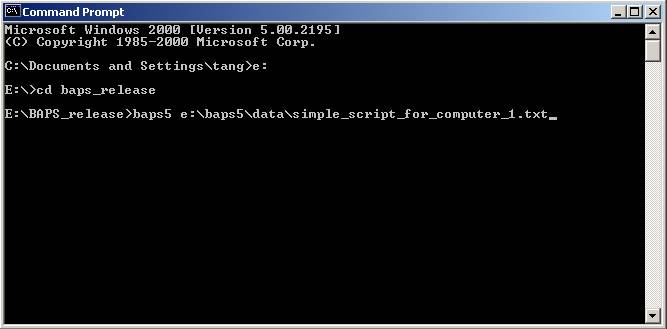
Figure 2: Start a sub-task with the script. Replace 'E:\BAPS_release'
with
the
installation directory of your own BAPS package. Also remember to put the full
path of your script file in the behind.
3) BAPS 5 will be evoked without
displaying the GUI and produce the result as the script required. (Figure 3)
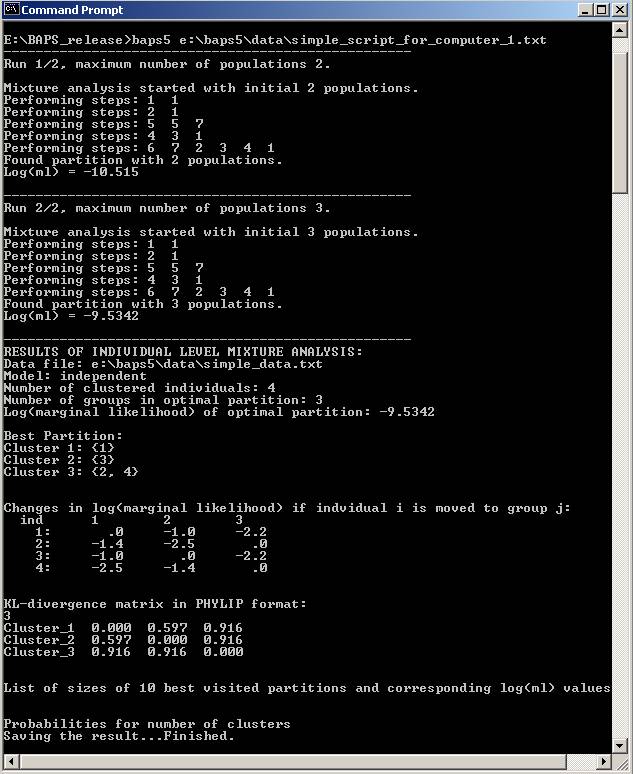
Figure 3: The partition result. The result has been
saved into the file as specified by the 'outputmat' command in the script.
4) Repeat the steps above on the second
computer to run the remaining subtasks. Here we show the second script to
obtain the mixture results for initial K = 4 and 5 (Figure 4).
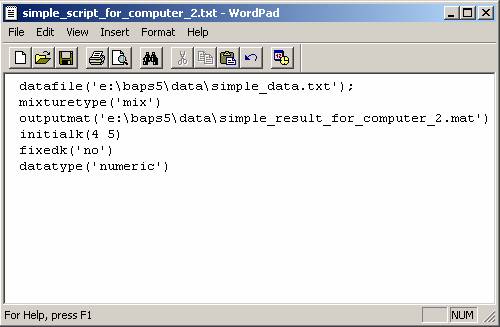
Figure 4: The
script for computer No.2. It starts a mixture analysis with initial K=4 and 5.
5) Collect all of the
result files into one computer and start the GUI version of BAPS 5.0. Click on
the 'Compare results' in the 'File' menu and load them into BAPS for comparison
(Figure 5). Here we loaded the two result files: simple_result_for_computer_1.mat
and simple_result_for_computer_2.mat. The comparing results will be shown in
the back window.
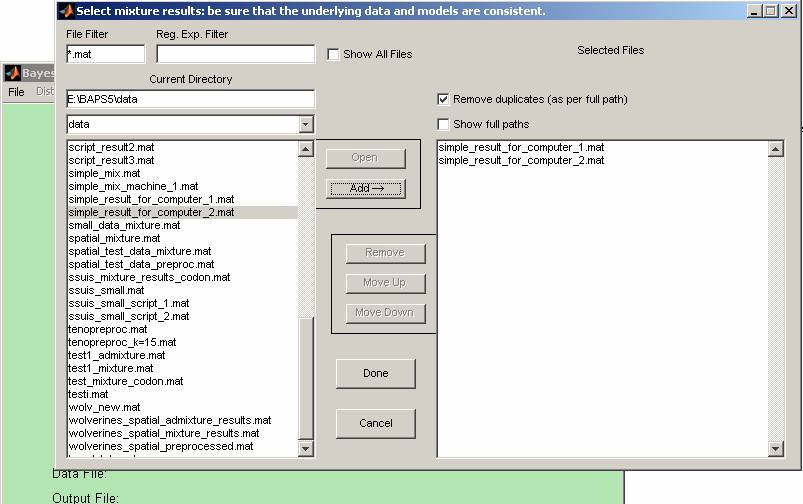
Figure 5: The partition file selection window
==================================
Q4: How to write a script file?
==================================
A4.1: Syntax
A script file
consists of several lines of commands. Each command line writes in a format as
commandname('parameter')
For example:
datafile('c:\data\data1.txt')
mixturetype('mix')
initialk(10)
...
NB!:
Non-numerical parameters are always enclosed in single quotation marks.
A4.2: Universal commands
-
Universal commands are mandatory and should be used irrespective of
mixture/admixture types.
datafile - Specify the full path of the data
datafile('c:\data\data1.txt')
mixturetype - Specify the mixture/admixture model
mixturetype('mix') - independent mixture model
mixturetype('linear_mix') - linear linkage model
mixturetype('codon_mix') - codon linkage model
mixturetype('spatial') - spatial mixture model
mixturetype('ad_mix') - admixture model
outputmat - Specify the result .mat file. The suffix of the result file
must be .mat
outputmat('c:\result\result1.mat')
A4.3: Mandatory commands for mixture
analysis
-
These commands should be used for any of the mixture models.
initialk - The initial number of clusters K
initialk(10 11 12)
initialk(
initialk(10) - if fixedk('yes') then use single value.
fixedk - Whether K is fixed during the analysis
fixedk('yes')
fixedk('no')
datatype - Specify the data type
datatype('numeric') - numeric baps data
datatype('sequence') - sequence baps data, for linkage model only
datatype('excel') - excel baps data, for linkage model only
datatype('matlab') - preprocessed baps data
datatype('genepop') - genepop data
A4.4: Optional commands for mixture
analysis
namefile - Specify the population name file
namefile('c:\data\data1_name.txt')
indexfile - Specify the population index file
indexfile('c:\data\data1_index.txt')
groups - Indicate clustering of groups instead of individuals
groups('yes')
groups('no')
groupname - Specify the group name file
groupname('c:\data\data1_groupname.txt')
coordinatefile - Specify the coordinate file, for spatial model only
when the data type is other than 'matlab'.
linkagemap - Specify the linkage map file, for linkage model only when
the data type is other than 'matlab' or 'excel'.
A4.5 Mandatory commands for
admixture analysis
clusters - Specify which of the clusters are analyzed
clusters(1 3 5) - cluster 1, 3 and 5 will be analyzed.
minsize - Specify the minimal size of population
minsize(5) - minimal size of population is 5 individuals
iters - Specify the number of iterations
iters(50)
refinds - Specify the number of reference individuals
refinds(10)
refiters - Specify the number of iterations for reference
individuals
refiters(20)
========================================================
Q5: Examples of script file
========================================================
Q5.1 Example 1 - BAPS data,
independent mixture model
datafile('r:\baps5\data\simple_data.txt');
mixturetype('mix')
outputmat('r:\baps5\data\simple_mix_1.mat')
initialk(2 3)
fixedk('no')
datatype('numeric')
Q5.2 Example 2 - Preprocessed data, independent
mixture model, fixed-k partition
datafile('e:\baps5\data\baps_data.mat');
mixturetype('mix')
outputmat('e:\baps5\data\baps_mix.mat')
initialk(10)
fixedk('yes')
datatype('matlab')
Q5.3 Example 3 - Excel data, codon linkage model
datafile('r:\baps5\data\simple_data.xls');
mixturetype('codon_mix')
outputmat('r:\baps5\data\simple_codon_mix.mat')
initialk(2 3)
fixedk('no')
datatype('excel')
Q5.4 Example 4 - BAPS data, spatial mixture model
datafile('r:\baps5\data\simple_data.txt');
mixturetype('spatial')
outputmat('r:\baps5\data\simple_spatial_mix.mat')
initialk(2 3)
fixedk('no')
datatype('numeric')
coordinatefile('r:\baps5\data\simple_cood.txt')
Q5.5 Example 5 - Admixture analysis on cluster 3 and 4
datafile('e:\baps5\data\ssuis_mixture_results_codon.mat')
mixturetype('ad_mix')
outputmat('e:\baps5\data\ssuis_small_script_2.mat')
clusters(3:4)
minsize(5)
iters(2)
refinds(3)
refiters(4)