mauve

- The Mauve home page is
http://asap.ahabs.wisc.edu/mauve/
- Here is a brief description of what Mauve does, copied from the Mauve home page:
Mauve is a system for efficiently constructing multiple genome alignments in
the presence of large-scale evolutionary events such as rearrangement and inversion.
Multiple genome alignment provides a basis for research into comparative genomics and the
study of evolutionary dynamics. Aligning whole genomes is a fundamentally different problem
than aligning short sequences.
Mauve has been developed with the idea that a multiple genome aligner should require
only modest computational resources. It employs algorithmic techniques that scale well in
the amount of sequence being aligned. For example, a pair of Y. pestis genomes can be
aligned in under a minute, while a group of 9 divergent Enterobacterial genomes can be
aligned in a few hours.
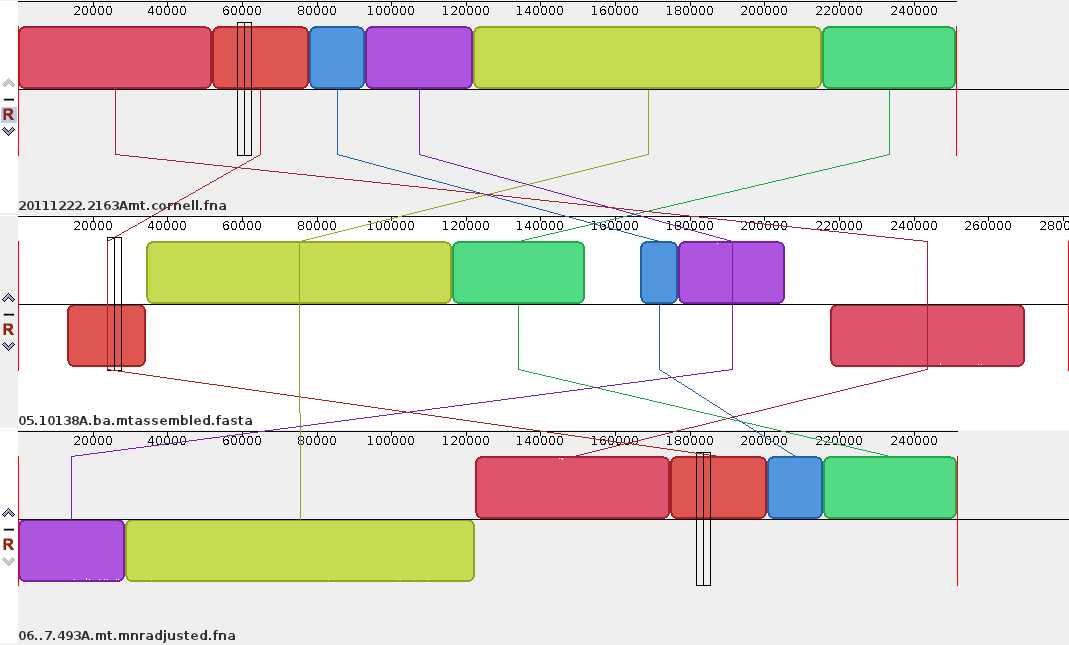
- Sample alignment of three carrot mitochondrial genomes